
An area where you can get a lot of data to analyze and apply many of the methods you are learning in this course, this NIH program. The NCI's, the National Cancer Institute's, TCGA, which is The Cancer Genome Atlas. 
For applications in personalized medicine and systems pharmacology. In this lecture, we are going to talk about integrating many of the components covered in the entire course. 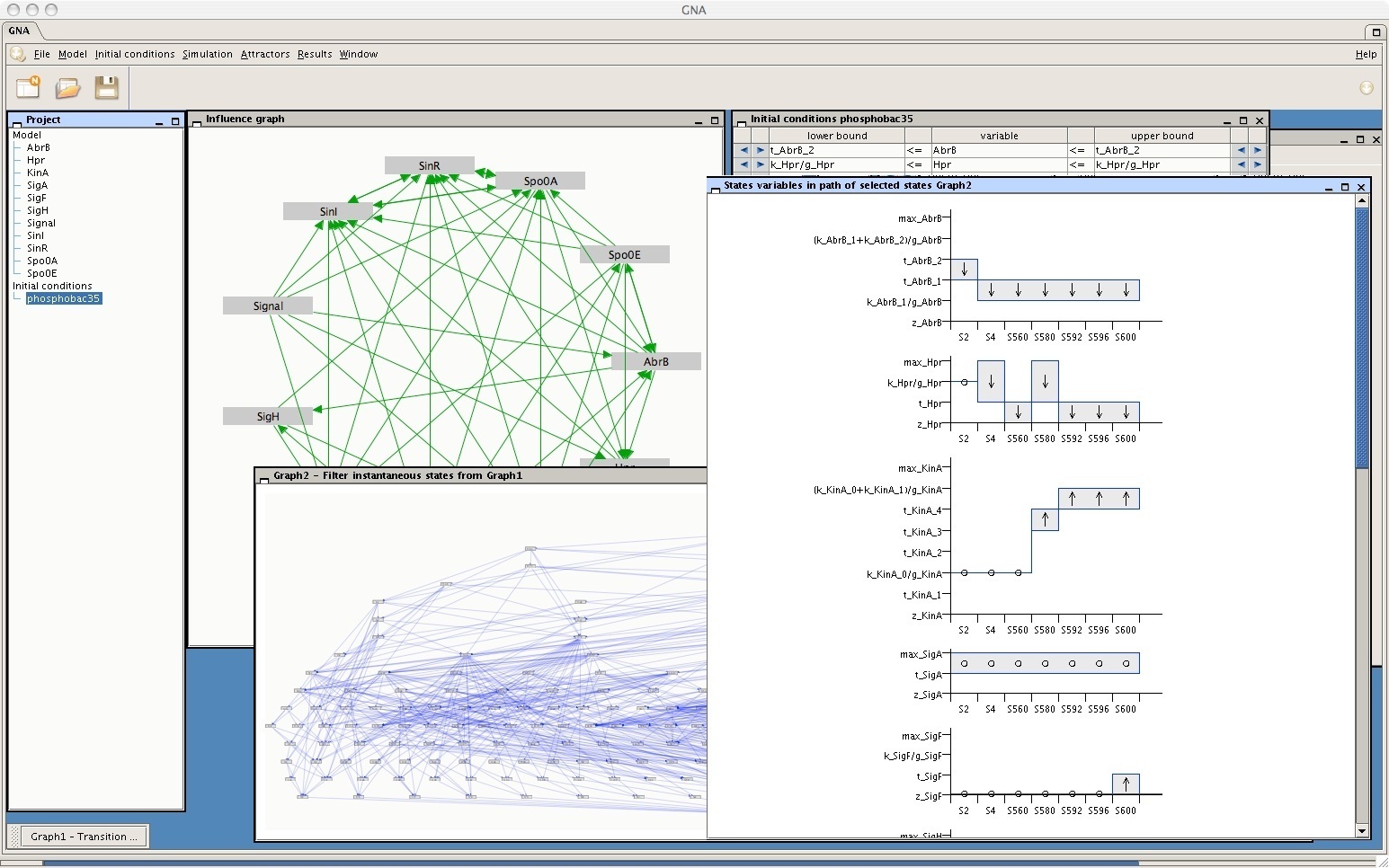
For those participants that do not work in the field, the course introduces the current research challenges faced in the field of computational systems biology. The ultimate aim of the course is to enable participants to utilize the methods presented in this course for analyzing their own data for their own projects. The course presents software tools developed by the Ma’ayan Laboratory () from the Icahn School of Medicine at Mount Sinai, but also other freely available data analysis and visualization tools. The course should be useful for researchers who encounter large datasets in their own research. The course is mostly appropriate for beginning graduate students and advanced undergraduates majoring in fields such as biology, math, physics, chemistry, computer science, biomedical and electrical engineering. 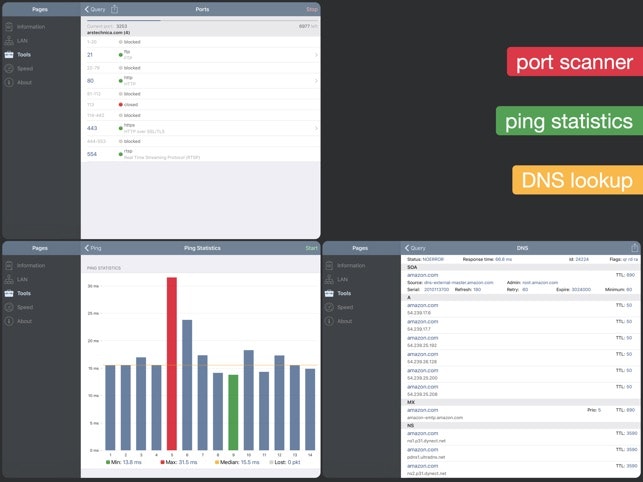
The course contains practical tutorials for using tools and setting up pipelines, but it also covers the mathematics behind the methods applied within the tools. The course covers methods to process raw data from genome-wide mRNA expression studies (microarrays and RNA-seq) including data normalization, differential expression, clustering, enrichment analysis and network construction. An introduction to data integration and statistical methods used in contemporary Systems Biology, Bioinformatics and Systems Pharmacology research.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
